Los genomas a través del árbol de la vida presentan un número de genes altamente variable que refleja diferencias en la complejidad biológica. La aparición de nuevos genes es un motor fundamental de la innovación evolutiva, ya que permite la adquisición de rasgos novedosos. La duplicación génica, propuesta por primera vez hace casi un siglo, constituye el principal mecanismo subyacente a la formación de nuevos genes. Esta revisión describe los mecanismos moleculares de la duplicación génica, así como los posibles destinos evolutivos y consecuencias funcionales de los genes duplicados, destacando su papel en la evolución de los organismos, la complejidad cerebral y las enfermedades humanas.
Emergence of New Genes and Gene Products: Mechanisms, Evolution and Functional Implications
1. Introduction
Large-scale DNA sequencing technologies that have been implemented during the past 20 years have enabled the analysis of the genome at an unprecedented scale across a wide range of species. These genomic studies have revealed that the number of genes varies enormously among organisms (Table I). The hypothetical Last Universal Common Ancestor (LUCA) of all three domains of life (Bacteria, Archaea, and Eukarya) has been proposed to hold around 2,600 protein-coding genes (Moody et al., 2024). These ancestral genes would have given rise to all the gene diversity that has been created on Earth ever since. Interestingly, the number of genes of modern free-living prokaryotes is very similar to that of LUCA, ranging from ~2,000 to ~4,000. By contrast, parasitic bacteria, such as Mycoplasma genitalium, typically hold fewer genes that enable them to fulfill only the most basic cellular processes as they depend fully on their host for many essential functions (Fraser et al., 1995). In comparison to prokaryotes, eukaryotes represent a leap in biological complexity, which is accompanied by an increase in gene number. For example, unicellular eukaryotes, such as yeast, have around 6,000 genes. A further expansion in gene repertoire appears to have been required upon the emergence of multicellularity. Thus, multicellular organisms, including nematodes, insects and mammals, such as humans, typically harbor genomes with around 14,000 to 22,000 protein-coding genes (Lander et al., 2001. Venter et al., 2001). These genes are necessary for coordinating the development and function of complex multicellular bodies, regulating key processes such as organogenesis, neural activity, immune responses, and behavior through sophisticated gene regulatory and cell signaling networks. Other organisms, typically plants, have an even larger number of genes. For example, the soybean Glycine max has a genome with over 46,000 protein-coding genes involved in a wide range of specialized functions, including nitrogen fixation through symbiosis with soil bacteria, complex developmental pathways, and sophisticated responses to environmental cues (Schmutz et al., 2010). In broad terms, the remarkable variation in gene number across species reflects the biological complexity and lifestyle demands of each organism.
Table I. Number of protein-coding genes of several organisms
| Organism | Number of genes |
|---|---|
| Mycoplasma genitalium (parasitic bacterium) | ~525 |
| Haemophilus influenzae (bacterium) | ~2,000 |
| Escherichia coli (bacterium) | ~4,400 |
| Saccharomyces cerevisiae (yeast) | ~6,000 |
| Drosophila melanogaster (fruit fly) | ~14,000 |
| Caenorhabditis elegans (nematode) | ~20,000 |
| Homo sapiens (human) | ~21,000 |
| Mus musculus (mouse) | ~22,000 |
| Zea mays (maize/corn) | ~32,000 |
| Glycine max (soybean) | ~46,000 |
Today, it is well-accepted that the generation of new genes is a key driver of evolutionary innovation, as it enables the emergence of new traits and adaptations. Species-specific or lineage-specific genes are found in many, if not all, organisms and serve as a blueprint for the acquisition of novel biological functions and the vast diversity of life. In this review, we provide a detailed overview of the major genomic event that gives rise to new genes in eukaryotes: gene duplication. Furthermore, we examine the evolutionary trajectories through which duplicated genes may become fixed in the population, as well as their relative contributions and functional significance in different species and evolutionary lineages, with a particular focus on animals. Next, we provide examples of how gene duplication has contributed to the emergence of important gene families present in multiple genomes. Finally, we focus on our own species to illustrate the “yin/yang” nature of gene duplications, as they have played a key role in the evolution of the human brain, a paradigmatic example of biological complexity, but they are also causally implicated in several human diseases. By exploring the emergence and diversification of new genes, we aim to gain a better understanding of the genomic players and evolutionary processes underlying biological complexity in living organisms.
The origin of new genes can be regarded as a paradigmatic example of biological emergence, in which novel properties arise from the reorganization, modification, and integration of pre-existing molecular components. New genes introduce functional elements that were not present in the ancestral genome and can give rise to novel biochemical activities, regulatory interactions, and phenotypic traits. Through their incorporation into gene regulatory and cellular networks, these genetic novelties contribute to increased biological complexity at higher levels of organization, including tissues, organs, organisms and their behavior. Similar to emergent phenomena in other scientific domains, these new properties cannot be easily predicted from individual components alone, highlighting the generation of new genes as a key molecular process for emergent complexity in evolution.
2. Duplication as a Mechanism to Form New Gene Structures
The identification of mechanisms underlying the generation of new genes has long been a subject of scientific interest. Over the years, researchers have uncovered a range of molecular processes that give rise to new genes. As early as in the 1930s, Haldane and Muller hypothesized that gene duplication could serve as a primary mechanism for the emergence of new genes. This idea was later developed by Ohno, who argued that a new duplicate copy of a gene could acquire a novel function and be retained in the genome by natural selection (reviewed in Long et al., 2003). These early notions that gene duplication provides a significant source of genetic innovation, phenotypic adaptation and biological complexity have since been broadly confirmed and refined, thanks to numerous molecular studies enabled by the genomics revolution. Indeed, gene duplication appears to be the most common way of creating new genes. New duplicate genes within a species, known as paralogs, have been shown to be abundant across all eukaryotic genomes sequenced to date and to have evolved essential functional roles (Lynch and Conery, 2000). Collectively, the entire set of paralogs within a species constitutes a gene family.
The duplication of genetic material that may lead to the emergence of new genes can occur via three main routes: DNA-based duplication, RNA-based duplication (retroposition) or a combination of both. These mechanisms will be described in more detail in the following sections.
2.1. The emergence of new genes through a DNA-based duplication process
DNA-based duplication mechanisms typically include the duplication of chromosomal segments containing whole genes or gene fragments (Betrán and Long, 2002). If they span from 1 kilobase (kb) to hundreds of kilobases in length, they are termed segmental duplications (SD). Duplications are typically the result of erroneous recombination (also called unequal crossover) processes during meiosis (either by non-allelic homologous recombination or by non-homologous end joining) and often involve repetitive DNA sequences that are scattered throughout the genome in high number (Liao et al., 2023). Of note, interspersed SDs are very abundant in humans and, according to the most recent sequencing analyses, represent up to 7% of our genome (Vollger et al., 2022). All these repetitive sequences provide regions of homology that may misalign during meiosis leading to aberrant recombination, and consequently to duplication of chromosomal segments in the resulting gametes (Figure 1).
Figure 1. Mechanism of DNA-based gene duplication
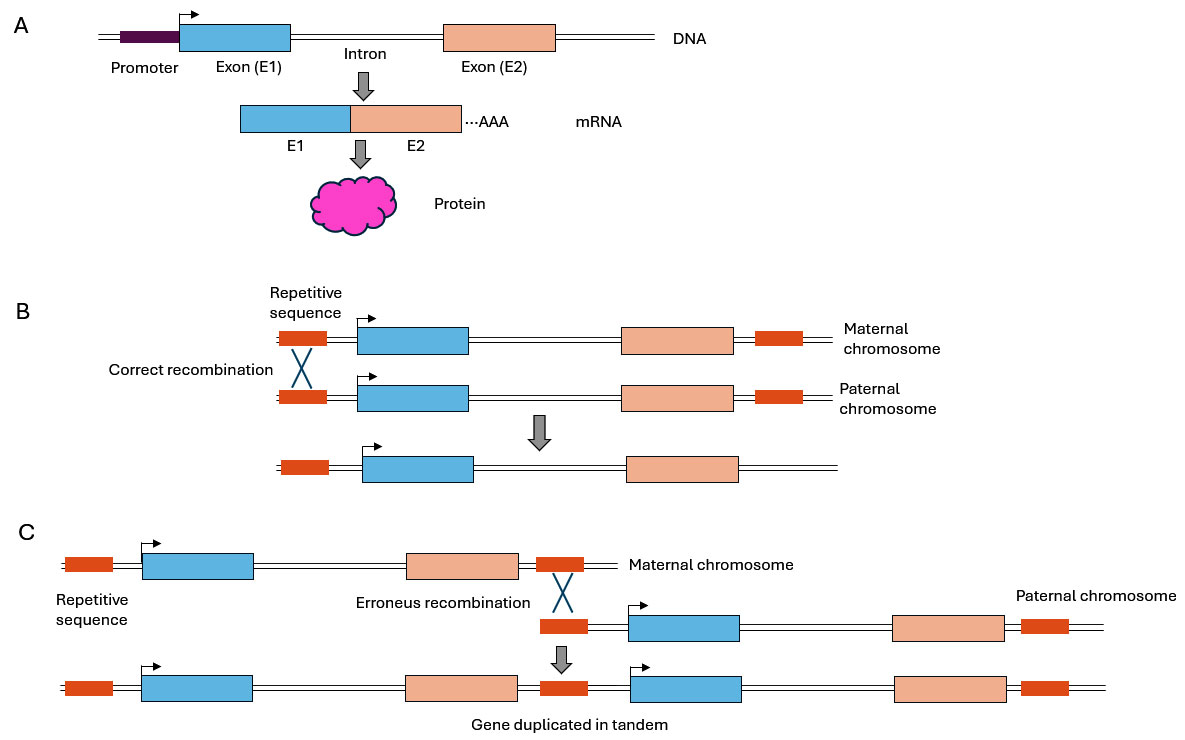
A) Schematic representation of the structure of a prototypical eukaryotic gene, showing the usual pattern of alternating of exons (protein coding regions, shown as light blue and light orange boxes) and introns (non-coding regions), as well as the transcription regulatory element known as promoter (purple) and the transcription initiation site (arrow). When a gene is active, it is transcribed into precursor mRNA, which then undergoes a series of processing and maturation steps before being translated into a functional protein.
B) Schematic mechanism of a correct meiotic recombination event (depicted as an X) between the maternal and paternal chromosomes during gametogenesis.
C) Schematic mechanism of an erroneous recombination event (unequal meiotic crossover), caused by misalignment of repetitive DNA sequences (represented as dark orange boxes), that results in the emergence of a tandem duplication.
DNA-based duplication mechanisms may also include duplication of whole genomes (WGD) through various polyploidization processes. During vertebrate evolution, two rounds of WGD have occurred, while other eukaryotes, particularly plants, have undergone multiple WGD events (reviewed in Kaessmann, 2010). Although a WGD event can lead to the duplication of thousands of genes, the vast majority of duplicates are usually lost over time. Nevertheless, a notable percentage of duplicated genes are preserved after WGDs: about 15% in teleost fishes after 350 million years, around 12% in yeast after 80 million years, and around 30% in plants of the genus Arabidopsis after 80 million years. These figures suggest that significant percentages of duplicated loci can be retained as so-called ohnologs (i.e. paralog genes originated from a WGD event) (reviewed in Long et al., 2013).
The rates of gene duplication are still a matter of debate. Using a molecular-clock approach based on synonymous nucleotide substitution rates between gene duplicates, Lynch and Conery suggested that gene duplication rates range from 0.0023 to 0.021 per gene per million years (Lynch and Conery, 2000). By contrast, a subsequent study employing a different approach (phylogenetic distribution of gene duplicates) obtained a much lower estimate (0.000391 to 0.000925 per gene per million years) (Zhou et al., 2008). Importantly, duplication rates are not constant over evolutionary time. For example, careful analyses in primates have identified a burst of gene duplication that occurred in the hominoid lineage, especially in humans and African apes, with a rate of more than 100 genes duplicated in the hominoid genome per 1 million years (Hahn et al, 2007a. Marques-Bonet et al., 2009). High rates (17 duplicated genes per 1 million years) have also been estimated in flies (Hahn et al., 2007b). In addition, the rate of duplication can remarkably differ across the genome. In fact, particularly elevated duplication rates of a small subset of gene families (0.016-0.03 per gene per million years) have been detected in diverse eukaryotic lineages. These families are associated with functions such as immunity, host defense, chemosensation, and reproduction, suggesting that gene copy number increase has significantly contributed to the adaptive evolution of these lineages (Emes et al. 2003. Demuth and Hahn 2009). Duplications of these genomic regions occur so frequently that they become polymorphic across individuals of the same species and contribute to differences in DNA content and gene number among them (Sebat et al., 2004). For example, any two humans are estimated to differ by approximately 5 megabases of DNA due to variations in the number of duplicated copies across specific genomic regions.
When the erroneous recombination event occurs within introns, exons can be exchanged between unrelated genes, through a process known as exon shuffling (Figure 2A). By rearranging exons, this process can create new combinations of protein domains, potentially leading to proteins with novel functions, and is thought to have played a crucial role in the evolution of multicellular organisms (metazoans), particularly in the development of complex, multidomain proteins involved in cell signaling and extracellular functions (Patthy, 2021).
Figure 2. Mechanism and physiological consequences of exon shuffling
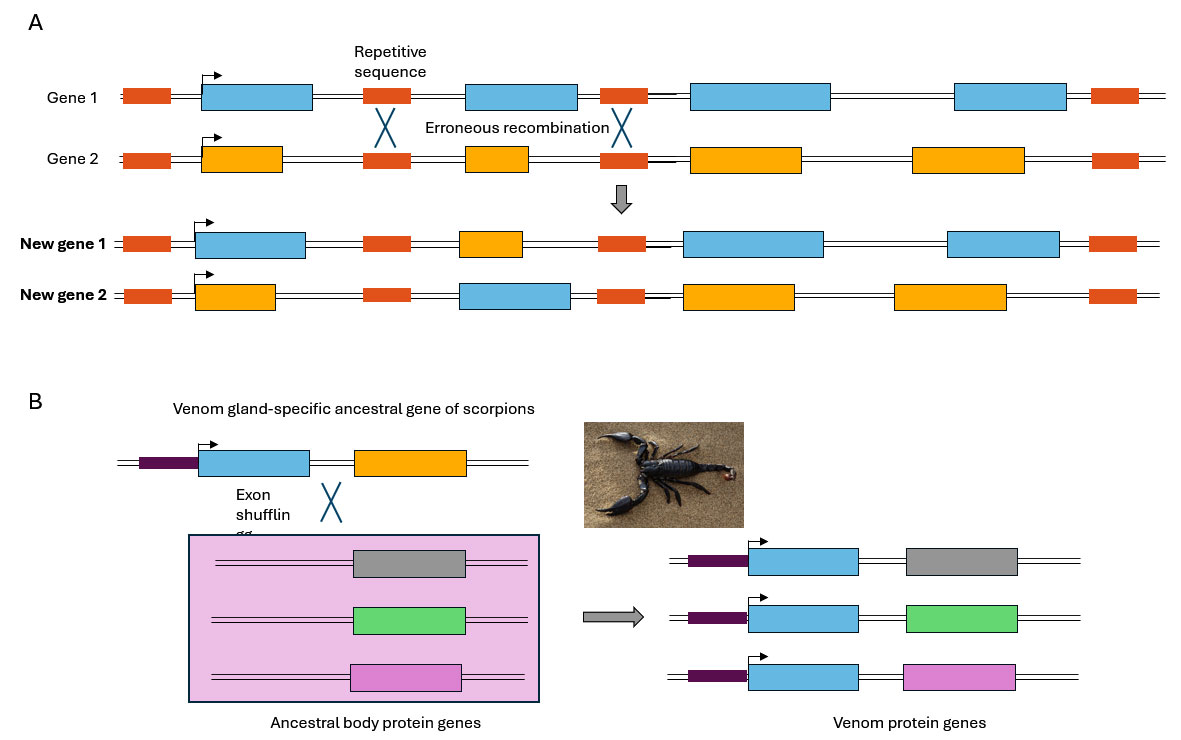
A) Schematic representation of the mechanism for the creation of a gene with a new sequence by exon shuffling. Erroneous recombination occurs between repetitive DNA sequences located in introns of two different genes.
B) The hypothetical evolutionary model for the origin of scorpion venom biodiversity by exon shuffling. Exon in blue encodes the secretory signal sequence. Modified from Wang et al., 2016.
An intriguing example of exon shuffling can be found in the evolutionary origin of scorpion venom toxins. It has been proposed that an ancestral gene expressed specifically in the venom gland, acted as a donor by providing two necessary regulatory elements for venom gland‐specific expression: a promoter and a secretory signal. A diversity of genes encoding body proteins would have served as sources of coding exons that contributed the sequence diversity of the toxins (Wang et al., 2016). These exon shuffling events are thought to have occurred multiple times throughout evolution in scorpion lineage, giving rise to an array of venom proteins with distinct specificities (Figure 2B).
2.2. The emergence of new genes through an RNA-based duplication process (retroposition)
In this mechanism, a messenger RNA (mRNA) that is transcribed from a given gene undergoes intron removal during maturation and is then reverse transcribed into a complementary DNA (cDNA) copy, which is inserted into the genome (Figure 3). This process requires an active reverse transcriptase activity in the cell, which, in mammals, is provided by LINE-1 (L1) retrotransposons present in high numbers in the genome (Feng et al., 1996. Esnault et al., 2000). The resulting retrocopied genes (retrogenes) only contain exon information, and lack parental introns.
Figure 3. Schematic mechanism of RNA-based gene duplication (retroposition)
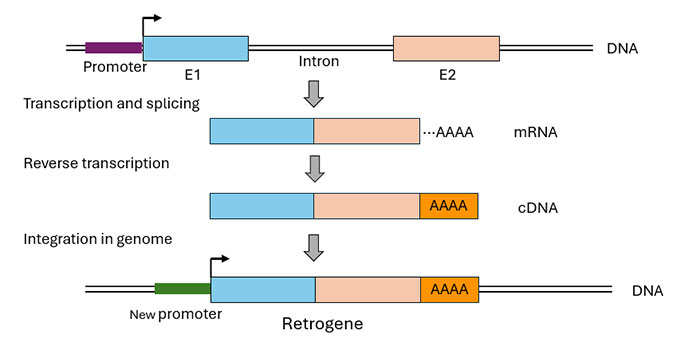
After transcription of a parental gene, the mRNA undergoes splicing and the intronless mature mRNA is reverse transcribed into a cDNA, which is then inserted in the genome.
In addition to introns, the newly integrated retrocopy also lacks promoter sequences, which are a necessary requirement for gene transcription and functionality. It is still unclear how retrocopies can obtain the regulatory sequences that allow them to be transcribed and thus, become functional retrogenes (an event that is more frequent than initially thought). Based on currently available evidence (reviewed in Kaessmann, 2010, Casola and Betrán, 2017), five different potential mechanisms have been proposed. First, new retrogenes can use the preexisting regulatory machinery of genes close to their insertion site. Second, retrogenes can recruit proximal CpG dinucleotide-enriched proto-promoter sequences not previously associated with other genes. Third, retrotransposons upstream of the insertion sites can provide retrogenes with regulatory potential. Fourth, retrogenes can directly inherit alternative promoters embedded in the parental transcripts that gave rise to them. Finally, retrogene promoters can evolve de novo through small substitutional changes under the influence of natural selection.
Numerous functional retrogenes have been detected in fruit fly genomes (Zhou et al., 2008). Similarly, thousands of retrocopies and over 100 functional retrogenes have been identified in the human genome (Vinckenbosch et al., 2006). In fact, because of the elevated activity of L1 retrotransposons, the overall rate of retroduplication has been high in most mammals with the exception of the oviparous group (e.g., platypus) (Kaessmann et al., 2009). In contrast, the genomes of oviparous mammals and birds contain very few retrocopies and appear to lack functional retrogenes, likely due to the absence of active retrotransposons in their genome (Hillier et al., 2004. Kaessmann et al., 2009).
Interestingly, retrogenes that eventually do become transcribed seem to be much more prone to evolve novel functions or specialized roles than gene copies arising from DNA-based duplication mechanisms (Vinckenbosch et al., 2006). PGK2 provides a well-characterized example of how retroposition can generate a functional retrogene with a specialized role. PGK2 arose by retrotransposition of PGK1, a gene in the X chromosome that encodes phosphoglycerate kinase 1, a key enzyme in glucose metabolism. During the retroposition process, a mature mRNA copy of PGK1 was reverse-transcribed and inserted into a new location on chromosome 6, giving rise to PGK2 (McCarrey, 1990). The original gene, PGK1, contains introns and is expressed ubiquitously. By contrast, the retrogene PGK2 lacks introns and is only expressed in testes, where it is thought to specifically support energy metabolism during spermatogenesis. Notably, many other autosomal retrogenes, including PDHA2 (encoding pyruvate dehydrogenase 2), and U2AF1L4 (encoding U2 Small Nuclear RNA Auxiliary Factor 1 Like 4), have also “moved out” of the X chromosome, indicating that there is strong positive selection for novel gene activity in male reproductive tissues during spermatogenesis (Long et al., 2003).
2.3 New genes arising from combined DNA- and RNA-based duplication mechanisms
Adding further complexity, new genes can also be generated through a combination of DNA-based and RNA-based duplication events, which would give rise to chimeric genes. A classic example is jingwei (jgw), a gene found in several Drosophila species (Figure 4). In a common ancestor of two African species of Drosophila, D. yakuba and D. teissieri, two single copy genes existed, yellow-emperor (ymp) and alcohol dehydrogenase (Adh). The yellow-emperor gene underwent a DNA-based duplication, resulting in two copies: ymp, which retained the original function, and yande (ynd), which was further involved in the origin of jingwei through and RNA-based duplication event. In this event, a retrocopy of the Adh gene became inserted downstream of the third exon of ynd. The resulting chimeric jgw gene contains the first three exons from ynd and the full retroposed Adh sequence, and encodes a fusion protein combining the sequences from both genes. The downstream exons of the original ynd, located after the Adh insertion, are no longer transcribed or translated, and have since accumulated mutations (Zhang et al., 2004).
Figure 4. Combination of DNA-based and RNA-based duplication mechanisms in the origin of the Drosophila gene jingwei
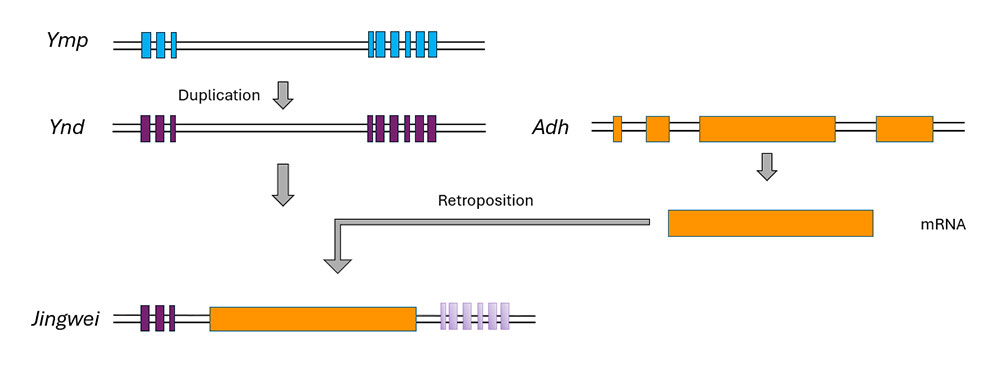
DNA-based duplication of ymp gave origin to ynd. Subsequent RNA-based retroposition of an Adh copy into ynd led to the origin of the jgw gene around 2 million years ago.
3. Evolutionary Fate of Duplicated Genes
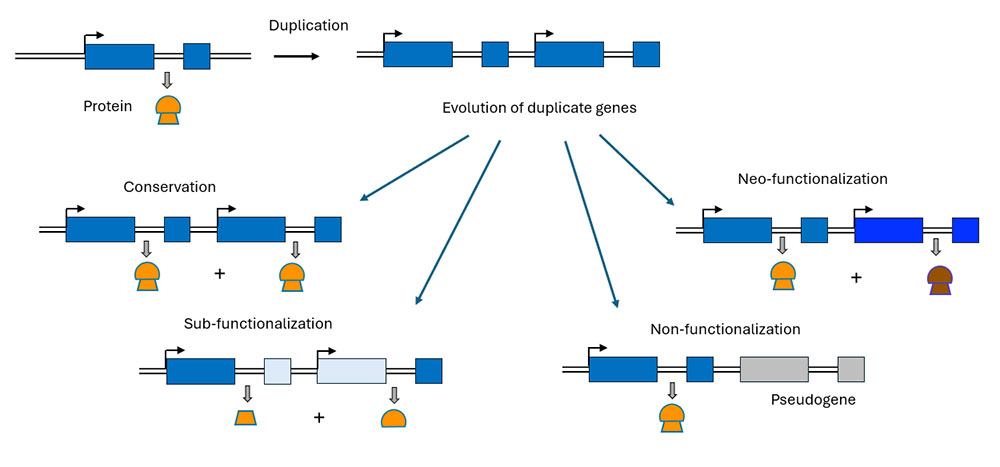
To understand how duplication events can lead to the emergence of new genes, we need to take into account the evolutionary processes involved (Xia et al., 2025). New gene emergence is thought to follow a microevolutionary process, whereby a novel protogene structure is initially generated in the germline (e.g. a gene duplication). This sequence needs to spread through the population until it becomes fixed within the entire species as a new gene. Evolutionary forces, particularly natural selection and genetic drift, will govern the fate of the protogene in the population, thus making protogene fixation a population genetic process that can result in different outcomes (Figure 5): non-functionalization, conservation, neo-functionalization or sub-functionalization (Prince and Pickett, 2002, Zhou and Wang, 2008).
3.1. Non-functionalization
After a gene duplication event, the two copies will initially be structurally and functionally identical and thus, functionally redundant. Because only one of the duplicates is required to maintain the ancestral gene’s function, the other copy becomes free from purifying selection and accumulates deleterious mutations, eventually becoming a pseudogene through a process called non-functionalization or pseudogenization (Ohno, 1973). In the human genome, approximately 12,000 to 14,000 pseudogenes have been identified (Pei et al., 2012), although some estimates suggest that this number could be close to 20,000 (Wright et al., 2016).
While most new gene copies will undergo this process, duplication events are also in the origin of many functional genes in present day genomes. For example, recent analyses of the human genome have revealed that at least 15% of human genes are indeed duplicates, and it has been estimated that around half of all duplicated vertebrate genes have been maintained (Nadeau andSankoff, 1997). As described below, other evolutionary processes that do not lead to pseudogenization account for the presence of duplicated, functional genes in the genomes of living species.
Figure 5. Different outcomes for a duplicated gene
3.2. Conservation
In many cases, the increased copy number after a duplication can lead to a higher expression level of the protein product encoded by the ancestral gene. If the effect of this gene dosage is beneficial (i.e. confers and adaptive advantage) there may be a strong selective pressure to preserve the duplicated copy (conservation) (Ohno, 1973). A prominent example is the AMY1 gene, which encodes salivary amylase, an enzyme that digests starch. Humans have a variable number of AMY1 gene copies, ranging from 2 to 15, which correlates with salivary amylase protein levels. AMY1 copy number variation (CNV) is thought to be an adaptation related to starch-containing diets, with populations consuming high-starch diets typically having more AMY1 gene copies. Earlier studies proposed that the AMY1 copy number increase was a relatively recent adaptation in response to agricultural diets, but recent high-resolution genomic analyses indicate that the AMY1 locus was already evolutionarily predisposed for adaptive expansion by the time agriculture spread (Figure 6). In fact, these analyses have revealed that the initial duplication event occurred as far back as 800,000 years ago, well before the out-of-Africa migrations of modern humans (Yilmaz et al., 2024). This gene locus represents one of the fastest-evolving duplicate repeat regions in the human genome.
Figure 6. Amylase (AMY1) gene duplication through evolution
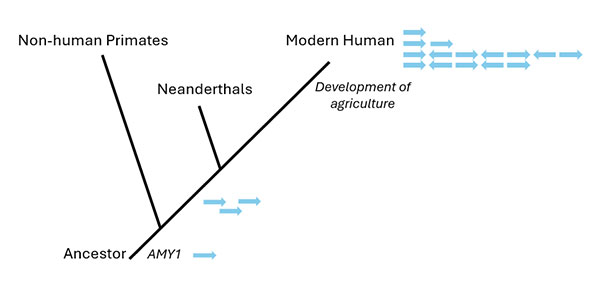
An initial duplication of the AMY1 gene is thought to have occurred ~800,000 years ago, even before the human-Neanderthal split, leading to the generation of three AMY1 genes. Hunter-gatherers already had variable AMY1 copy numbers as early as 45,000 years ago, followed by a remarkable increase in the AMY1 copy number in the genomes of farmers over the past ~8000 years. Modified from Yilmaz et al., 2024.
A curious example of gene dosage-related conservation of a duplicate copy was reported by Parker et al. (2009). They found that a retrogene derived from a growth factor gene (fgf4) involved in bone development is responsible for the short-legged phenotype characteristic of several common dog breeds, including the dachshund (Figure 7). Remarkably, the coding sequence of the retrocopy is identical to that of its parental gene and thus, the phenotypic impact of the duplication seems to be a rather direct consequence of the associated gene dosage (i.e., increased FGF4 expression during bone development). This example also clearly illustrates how gene duplication can immediately lead to phenotypic innovation (in this case a new morphological trait) under human-induced artificial selection imposed by breeding.
Figure 7. Image of a short-legged dachshund

3.3. Neo-functionalization
Following a duplication event, one gene copy can maintain the original function, while the other is free to accumulate mutations that may lead to novel functions, a process known as neo-functionalization (Kimura, 1983). According to this model, gene redundancy would allow for relaxed purifying selection on the duplicate, which would be free to accumulate mutations that would be deleterious in a single-copy context. Under certain conditions, such as environmental change, dietary shifts, sexual selection, etc., some of the mutant alleles that encode a new function could confer advantageous traits, and therefore be preserved by positive selection. This adaptive evolution model is supported by several case studies and theoretical works showing that the evolution of recently created genes involves rapid changes in both gene structure and function (reviewed in Long et al., 2003).
An illustrative case of neo-functionalization is provided by the duplication of a pancreatic ribonuclease gene (RNASE1) in some species of leaf-eating monkeys around 4.2 million years ago. Under the influence of strong positive selection (posed by the demand for increased enzymatic activity in a microbially rich environment, such as the digestive system following a dietary shift) RNASE1B, a duplicate of the ancestral RNASE1 gene, rapidly adapted to digest nutrients from bacteria in the foregut of an African colobine monkey (Zhang et al., 2002). Remarkably, both the duplication and subsequent neo-functionalization of this gene occurred independently in a very similar manner in an Asian colobine (Schienman et al., 2006. Zhang 2006). In both cases, the newly evolved gene, RNASE1B, underwent rapid amino acid changes driven by Darwinian selection. A 3% change in amino acid sequence led to an increase in the negative charge and an altered optimal pH of the RNASE1B protein in both monkey species. As a result of these changes, which are essential for its newly evolved digestive function, RNASE1B lost the non-digestive activity (the ability to degrade double-stranded RNA), that is the hallmark function of the ancestral RNASE1. In addition to exemplifying functional specialization and relaxation of purifying selection, the convergent evolution of RNASE1B in both African and Asian colobines highlights that, although mutations are generated randomly, natural selection can shape similar outcomes at the molecular level when faced with similar ecological pressures.
An extreme case of neo-functionalization has been unveiled in a mouse retrocopy of a ribosomal protein gene (Rps23), of which there are hundreds of copies in mammalian genomes and that usually represent nonfunctional retropseudogenes, consistent with the idea that duplication of these genes is usually redundant and/or is subject to dosage balance constraints. Yet an Rps23 retrocopy evolved a completely new function, not by changes in the protein-coding sequence, but because it is transcribed from the reverse strand of the DNA sequence, and because it incorporated sequences flanking its insertion site as new exons (Zhang et al. 2009). This gave rise to a new protein (completely unrelated to that encoded by its parental gene), which had profound functional implications because it conferred increased resistance in mice against the formation of Alzheimer causing amyloid plaques.
3.4. Sub-functionalization
A gene may have multiple, often pleiotropic, functions. It has been proposed that the multiple functions of an ancestral gene may be partitioned after duplication between the two daughter copies, so that together, they retain the full ancestral function. This process was termed sub-functionalization, and may be shaped by natural selection or involve purely neutral processes (Force et al., 1999). This model proposes that, after duplication, the two gene copies accumulate complementary loss-of-function mutations in independent sub-functions, such that both genes must work together to perform the ancestral task (reviewed in Prince and Pickett, 2002).
Globin genes are a prime example of sub-functionalization, where duplicated genes complement the functions of the ancestral gene, rather than one paralog acquiring a new function. Approximately 450–500 million years ago, a single ancestral globin gene duplicated, giving rise to separate α‑globin and β‑globin gene families (Hoffmann et al., 2010), which underwent sequence and regulatory divergence. In humans, adult hemoglobin is a heterotetramer composed of two α‑chains and two β‑chains (α₂β₂) (Figure 8). Importantly, neither α nor β chains can form functional hemoglobin alone, as tetramers composed solely of one chain are nonfunctional. This interdependence reflects classic sub-functionalization: each paralog renounces part of the ancestral function, and together they restore full activity (Aguileta et al., 2004). In addition, other globin paralogs show specific developmental expression patterns during embryonic and fetal stages, thus reflecting sub-functionalization through temporal regulation (Hoffmann et al., 2010).
Figure 8. Structure of human adult hemoglobin
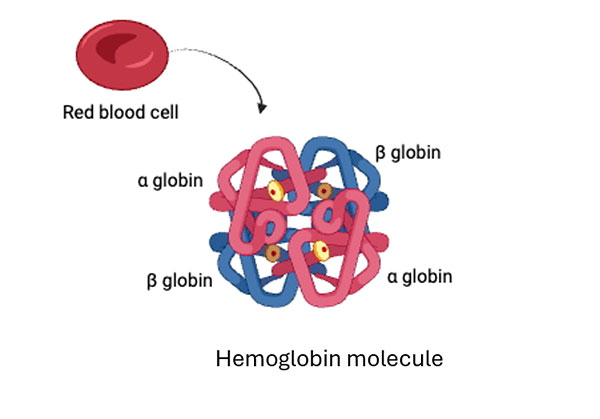
It is a heterotetramer composed of two α‑globin chains and two β‑globin chains (α₂β₂). Created in https://BioRender.com
Collectively, these different evolutionary outcomes illustrate how gene duplication can drive functional innovation and contribute to biological diversity. While non-functionalization may lead to gene loss in some cases, the retention and divergence of duplicates through conservation, neo-functionalization, or sub-functionalization generates new gene functions, modifies gene regulatory networks, and allows for the specialization of molecular pathways. In this way, the fate of duplicated genes is not simply a matter of molecular redundancy, but a fundamental process through which genomes acquire novel functional capabilities, ultimately shaping the complexity and adaptability of living organisms.
4. Role of Duplication in the Evolutionary History of Gene Families
Present day genomes contain gene families, that is, clusters of paralogous genes that share a common ancestral gene and arose through a series of duplication events. These paralogs frequently retain related biological functions. In humans, gene families comprise the largest proportion of the protein-coding sequences (Dornburg et al., 2022). The origin and evolutionary history of many gene families reflect a complex interplay of gene duplication, natural selection and genetic drift, and illustrate the functional diversification that emerges from the different fates of the duplicated copies (non-functionalization, conservation or neo/sub-functionalization). In the following section we provide two examples of gene families that highlight the diverse evolutionary outcomes experienced by paralogs.
4.1 The E2F family of transcription factors
E2F transcription factors are key regulators of the cell cycle by activating or repressing the expression of genes involved in DNA replication, mitosis, DNA repair, differentiation, development, and apoptosis (Iglesias-Ara and Zubiaga, 2015. Kent and Leone, 2019). In mammals, the E2F family includes eight members (E2F1–E2F8), which collectively contribute to their broad spectrum of cellular functions. Importantly, the E2F family is absent in prokaryotes, but is quite well conserved across eukaryotic lineages, with homologues identified in both unicellular and multicellular organisms (Templeton et al 2004. Iyer et al., 2008. Cao et al, 2010). Their widespread presence suggests that the E2F-dependent regulatory axis emerged early in eukaryotic evolution, likely playing a basic role in regulating cell division and maintaining genome stability.
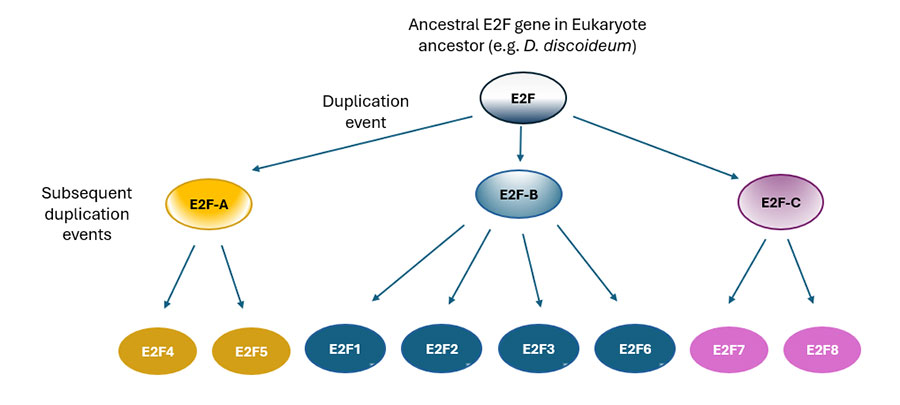
Phylogenetic comparisons indicate that a key gene duplication event occurred before the divergence of placozoans and bilaterians, giving rise to two main E2F subgroups, E2F-A and E2F-B (Cao et al, 2010). These two genes would represent a classic example of neo-functionalization, where one copy (E2F-A) retained the original transcriptional-repressive roles of the ancestral gene, while the other (E2F-B) evolved transcription activating functions. A third subgroup, represented by a hypothetical E2F-C, is thought to have emerged from an earlier duplication. In subsequent duplication events, E2F-A further diversified into E2F4 and E2F5, and E2F-B into E2F1, E2F2, E2F3 and E2F6, while E2F-C gave rise to E2F7 and E2F8 (Figure 9).
This view of the evolutionary history of the E2F family is further supported by molecular evolution analyses of present day E2F genes. Thus, reflecting the above-mentioned neo-functionalization of E2F duplicates, E2F4 and E2F5 (derived from E2F-A) show high similarity to the ancestral E2F sequence, evolve slowly, and are subject to strong purifying selection, consistent with maintenance of ancestral roles (Cao et al., 2010), while E2F1–3 (derived from E2F-B) exhibit faster evolution and weaker purifying selection, suggesting they have acquired novel, lineage-specific roles. On the other hand, E2F7 and E2F8 (derived from E2F-C) are structurally and functionally distinct form the other E2Fs: they contain two DNA-binding domains instead of one, and function as transcriptional repressors through mechanisms that differ from those of E2F4/5 (Cao et al., 2010). Further underscoring their evolutionary novelty, vertebrate E2F7/8 are known to play specialized roles in trophoblast biology and embryogenesis.
Together, the evolution of the E2F gene family illustrates how gene duplication, followed by functional divergence, can drive the emergence of regulatory complexity. The preservation of ancestral functions among E2F4/5 and the acquisition of new, activating or repressive roles by E2F1, 2, 3 and 6 and E2F7/8 reflect the evolutionary flexibility of transcription factor networks. This diversification likely enabled a more fine-tuned control over cell proliferation, differentiation, and development, critical processes in the transition to multicellular life. By expanding and specializing, the E2F family exemplifies how transcriptional regulators evolve in concert with increasing organismal complexity.
Figure 9. Proposed generation and functional diversification of E2F family members
4.2 The olfactory receptor family
The olfactory receptor (OR) gene family, the largest family in vertebrates resulting from extensive gene duplication of an ancestral gene, is a classic example of a combination of non-functionalization and neo-functionalization. In humans, there are over 800 genes in this family, approximately half of which are pseudogenes (Figure 10). In other species, like rats or horses, the number of OR genes is even higher, reflecting their higher use of the olfactory sense. The extensively duplicated olfactory receptor genes evolved new odorant specificities, which allowed for enhanced sensory capabilities and increased ecological adaptability (Olender et al., 2020). At the same time, there has been a massive wave of OR pseudogenization. In the case of primates, this has been associated with a reduced dependence on the sense of smell and compensatory reliance on other senses (e.g., vision).
Figure 10. Number of olfactory receptor genes in various vertebrate genomes
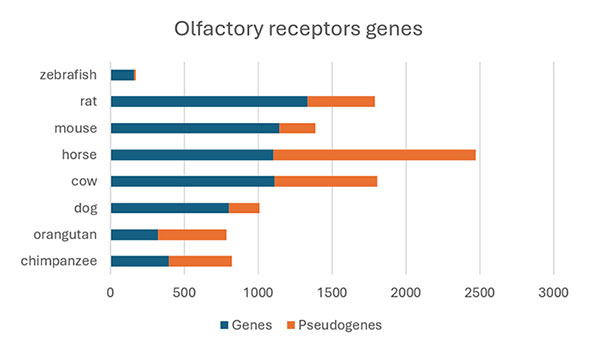
Shown is the number of functional and non-functional OR genes in each species. Modified from Olender et al., 2020.
5. Role of Duplication in the Evolution of Human Brain
The human brain is characterized by a particularly expanded cerebral cortex that is largely responsible for the cognitive capacities that distinguish our species. It represents a paradigmatic example of complexity, and the role that genetic determinants have played in human brain evolution has been a focus of intense research. A common approach to tackle this question has been to compare the genomes of humans and other closely related primates, such as the chimpanzees, searching for human-specific genes with a function in neurodevelopmental processes (Fair and Pollen, 2023). Interestingly, several of these genes (including SRGAP2, ARHGAP11 and NOTCH2NL, described below) have turned out to reside in segmental duplications (SDs), and to have undergone human-specific duplication events that contribute to the increased volume and complexity of human cortex (Dennis and Eichler, 2016. Suzuki and Vanderhaeghen, 2018, Soto et al., 2023).
The first described example of a set of neurodevelopment-related genes specifically duplicated in human was the SRGAP2 gene family. Besides the ancestral SRGAP2A copy, our genome contains three human-specific paralogs (SRGAP2B, C and D) that emerged as a result of incomplete duplication events, and thus, encode truncated proteins (Dennis and Eichler, 2016. Suzuki and Vanderhaeghen, 2018, Soto et al., 2023). These shorter proteins, most prominently SRGAP2C, can heterodimerize with SRGAP2A and interfere with its function (Charrier et al., 2012. Dennis et al., 2012). Through this dominant-negative effect, SRGAP2C delays neuronal maturation and increases the density of dendritic spines, thus influencing cortical connectivity.
Another human-specific incomplete duplication event, which generated a novel, truncated copy of ARHGAP11A, termed ARHGAP11B, was also identified (Antonacci et al., 2014). Because of subsequent genetic changes in ARHGAP11B, the encoded protein bears a carboxy-terminal end different from ARHGAP11A, and seems to have evolved a completely novel function (Soto et al., 2023), thus providing a clear example of neo-functionalization. Importantly, ARHGAP11B has been experimentally shown to promote basal progenitor amplification and expansion of the neocortex (Florio et al., 2015).
More recently, four human-specific paralogs of the NOTCH2 gene, termed NOTCH2NLA, B, C and R, have been identified and characterized (Fiddes et al., 2018. Suzuki et al., 2018. Bizzotto and Walsh, 2018). While NOTCH2NLR encodes a highly unstable protein that is likely non-functional, the proteins encoded by NOTCH2NLA, B, and C have been experimentally shown to increase the proliferation of neural progenitor cells and to delay neuronal differentiation, thus contributing to boost cortical neurogenesis (Fiddes et al., 2018. Suzuki et al., 2018). These examples underscore the role of duplication as a fundamental engine for the emergence of new functions and the increase in organismal complexity.
6. Duplications in the Human Genome: A Double-Edged Sword
As described in the sections above, duplication of genomic regions constitutes a major driving force in the evolution of species, including ours. It must be noted, however, that the immediate consequences of some duplication events may be deleterious for an individual organism. In fact, gene duplications are a well-recognized cause of human disease (Conrad and Antonarakis, 2007). Because of their highly homologous and interspersed nature, SDs provide a substrate for chromosomal misalignment and erroneous recombination during meiosis, thus posing a risk for chromosomal rearrangements (deletions, duplications, inversions or translocations) caused by the process of non-allelic homologous recombination (Emanuel and Shaikh, 2001). In particular, SDs are common mediators of the so-called recurrent genomic rearrangements (i.e., those that occur independently in multiple individuals, share a common size and show similar breakpoints). These genomic defects cause a variety of human diseases, where the inappropriate dosage of one or more genes has a pathological effect.
For example, SDs mediate a recurrent genomic rearrangement on the short arm of chromosome 17 leading to the duplication of the single gene, PMP22 (Figure 11A). Duplication of this gene is the most common cause of the Charcot-Marie-Tooth type IA (CMT1A, OMIM #118220) syndrome (Lupsky, et al., 1992), where the production of too much PMP22 protein causes breakdown of the protective myelin sheath around the nerves, resulting in peripheral nerve damage and muscle weakness in affected individuals.
Figure 11. Examples of the role of segmental duplications in human disease
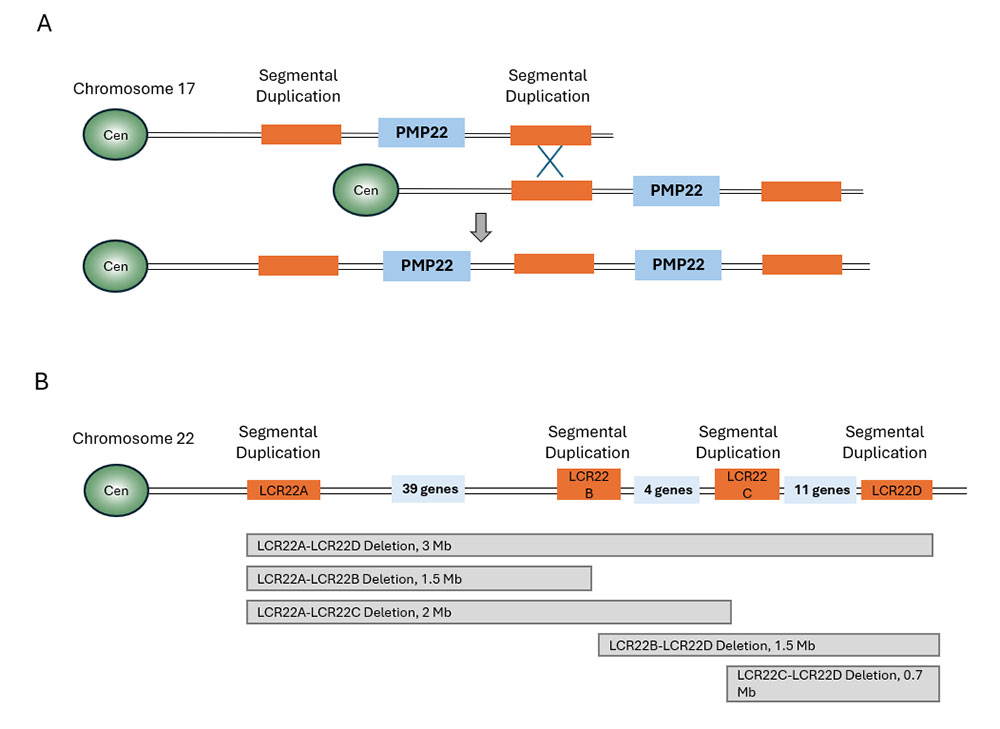
A. Schematic representation of the human chromosome 17 segment where the PMP22 gene is located. Shown is the erroneous meiotic non-allelic recombination event between segmental duplications in homologous chromosomes that gives rise to a duplicated PMP22 gene, causing CMT1A syndrome.
B. Schematic representation of the human chromosome 22 segment containing four SDs that are involved in erroneous recombination events leading to 22q11.2 deletion syndrome. The number of genes located between the SDs is indicated in the blue boxes. Grey boxes illustrate the different deletions that can be found in patients with this syndrome. Cen, centromere. Modified from McDonald-McGinn et al, 2015.
The importance of SDs as triggers of recurrent genomic rearrangements is further evidenced by their causative role in the most common microdeletion disorder in humans, the 22q11.2 deletion syndrome (OMIM #192430), which occurs in approximately 1 in every 1000 pregnancies. The 22q11.2 region harbours a set of 8 highly identical SDs, usually referred to as LCR22A to H, which makes this region one of the most complex and unstable loci in our genome (Vervoort and Vermeesch, 2022). Different non-allelic homologous recombination events between some of these SDs (LCR22A to D) may result in a variety of genomic rearrangements, including deletions and duplications of variable size, which leads to improper dosage of multiple genes (Figure 11B). The most common alteration is a 3 Mb deletion caused by NAHR between LCR22A and LCR22D, which affects about 50 protein-coding genes (McDonald-McGinn et al., 2015). Consistent with the complex nature of the underlying genomic defect, individuals affected by the 22q11.2 deletion syndrome show a highly heterogeneous set of clinical manifestations that often include heart and palate defects, immune deficiency and learning delay, among others (McDonald-McGinn et al., 2015). Although less frequent than the 22q11.2 deletion syndrome, the reciprocal 22q11.2 duplication syndrome (OMIM #608363) has also been described.
Finally, the recurrent 1q21.1 duplication and deletion syndromes, whose clinical features include brain size anomalies (macrocephaly in the case of 1q21.1 duplications, and microcephaly in the case of 1q21.1 deletions), have been causally linked to the NOTCH2 gene paralogs NOTCH2NLA, B, and C genes (Fiddes et al., 2018). These observations further reinforce the role of duplicated NOTCH2NL paralogs in modulating human brain size, but also illustrate the yin/yang nature of these gene duplications.
7. Conclusions
The examples of gene duplication discussed in this review underscore its role as a fundamental evolutionary mechanism that simultaneously drives innovation and carries inherent risks. By generating the raw material for genetic novelty, duplication events have shaped the complexity of genomes and have contributed to the emergence of the remarkable biological diversity observed on Earth. At the same time, these events can predispose genomes to instability and thus, contribute to disease.
A better understanding of the balance between these beneficial and deleterious outcomes will not only offer insights into the mechanisms of evolution, but also establish a valuable framework for leveraging gene duplication processes in biomedical research and therapeutic applications.
8. Glosary of Terms
Allele Each one of the two or more alternative forms of a gene. Different alleles determine the specific expression of the trait encoded by the gene in an individual.
Autosomal Pertaining to any trait encoded in chromosomes that are not involved in sex determination.
Bilaterian A member of the Bilateria, a major group of animals characterized by bilateral symmetry (i.e. having a distinct front, back, top, and bottom) and triploblastic development (i.e. possessing three germ layers: ectoderm, mesoderm, and endoderm).
Chimeric Gene A gene bearing a hybrid sequence resulting from the combination of two previously independent sequence segments.
Chromosomal (or Genomic) Rearrangements A broad class of large-scale structural alterations that modify the physical architecture of the genome. These alterations often result from errors during recombination or DNA repair processes and include:
- Deletions: Loss of a DNA segment, resulting in the removal of genetic material.
- Duplications: Repetition of a specific DNA segment, leading to an increase in genetic material.
- Inversions: A chromosome segment is excised and reintegrated in the same location, but in reverse orientation.
- Translocations: The relocation of a chromosome segment to a new physical location in a different chromosome.
Copy Number Variation (CNV) A type of structural genomic variation where segments of the genome are repeated, and the number of repeats varies among individuals. CNVs may significantly impact gene dosage and phenotypic diversity.
Deleterious Mutation A DNA sequence modification that negatively impacts the operational efficiency, stability, or survival probability of the organism.
Dominant-Negative Effect A phenomenon whereby a mutant allele actively interferes with or inhibits the function of its wild-type counterpart.
Eukaryote An single-celled or multicellular organism whose genetic material (DNA) is organized in the form of chromosomes and enclosed within a distinct nucleus. Besides the nucleus, eukaryotes possess other membrane-bound organelles (e.g., mitochondria, endoplasmic reticulum) and cells divide by a process called mitosis.
Exon A segment of a gene that remains in the mature mRNA after splicing.
Fusion protein A polypeptide encoded by a chimeric gene that contains amino acid sequences originally belonging to separate proteins.
Gene Dosage The number of copies of a specific gene present in the genome of an organism.
Genetic Drift A stochastic process whereby the frequency of specific traits or alleles in a population fluctuates due to random sampling rather than selection.
Genome The complete set of genetic material in a cell or organism, usually DNA (or RNA in some viruses), containing all the information needed to build and maintain that organism.
Germline The specialized cellular lineage that gives rise to gametes (e.g. egg and sperm cells), and thus mediates the transmission of biological information to subsequent generations.
Heterodimer A functional complex formed by the association of two non-identical molecular subunits.
Intron A sequence within a gene that is transcribed into precursor mRNA, but is subsequently removed during RNA processing before RNA is used to translate into protein.
Locus The specific, fixed physical position occupied by a gene within the chromosome.
Loss-of-function Mutation A mutation that results in reduced or abolished protein function.
Microevolution Progressive change in the allele frequencies within a population's gene pool over successive generations.
Natural Selection / Darwinian Selection An evolutionary process in which organisms with genes that are better suited to their environment are more likely to survive and reproduce, passing those advantageous genes to the next generation.
Non-Allelic Homologous Recombination (NAHR) A recombination process whereby similar DNA sequences that are not in the same position on chromosomes are incorrectly exchanged, often triggering genomic rearrangements, such as deletions, duplications or translocations.
Non-Homologous End Joining A rapid DNA repair mechanism that fuses broken molecular ends without using a reference template sequence, prioritizing structural continuity over information fidelity.
Ohnologs A particular type of paralogs resulting from a singular, large-scale doubling event of the entire genome (whole genome duplication).
Paralogs Evolutionary related genes that are generated by a duplication event within a single genome, often developing new or specialized functions.
Phenotypic Adaptation A modification in the observable physical or behavioral characteristics of an individual that enhances its functional success in a particular environment.
Phylogenetic Relating to the evolutionary history and relationships of a species or group of organisms, showing how they have diverged and diversified over time.
Placozoan A member of the Placozoa, a basal group of non-bilaterian, multicellular animals that display very simple structure, lacking organs, specialized digestive tracts or nervous systems. This group provides crucial information about early metazoan evolution.
Pleiotropic A property whereby a single gene exerts influence over multiple, seemingly independent systemic functions or traits.
Polymorphic The presence of multiple distinct allelic variants of a single gene.
Polyploidization A process whereby an organism acquires one or more extra copies of its complete set of chromosomes.
Prokaryote A microscopic single-celled organism that lacks a distinct nucleus and other specialized membrane-bound organelles. Prokaryotes include bacteria and archaea.
Proto-promoter A region of DNA that was originally inactive but acquires the ability to drive the transcription of a gene.
Protogene A genome sequence in the process of transitioning from being non-functional to becoming an active, transcribable gene.
Pseudogene A non-functional copy of a gene that bears sequence and structural similarity to the ancestral gene, but lacks the capacity to be expressed and often accumulates mutations.
Purifying Selection An evolutionary “corrective” process that removes detrimental genome variations to preserve the integrity of the original sequence.
Recombination The process by which genetic material is rearranged to form new combinations of alleles. This occurs naturally during meiosis via crossing-over between homologous chromosomes, and is a major force driving genetic variation in offspring.
Retrogene A functional gene created through retroposition. Unlike the ancestral gene copy, a retrogene typically lacks introns and the original promoter, often relying on "hitchhiking" a nearby promoter to be expressed.
Retroposition A process of gene duplication whereby a mature mRNA copy of a gene, lacking introns, is converted back into DNA and inserted into a new position in the genome.
Retropseudogene A non-functional copy of a gene generated by retroposition. In addition to lacking introns, they usually contain mutations that prevent their expression.
Retrotransposons Mobile elements that copy themselves by converting their RNA sequence back into DNA through a process called reverse transcription, then inserted the new DNA copy into the genome.
Reverse Transcription The biochemical process of synthesizing DNA from an RNA template, catalyzed by the enzyme reverse transcriptase. This process is a hallmark of retroviruses and retrotransposons.
Secretory Signal (Signal Peptide) A short peptide sequence (typically 16–30 amino acids long) present at the N-terminus of newly synthesized proteins that directs them to the secretory pathway (endoplasmic reticulum) for transport out of the cell or to specific organelles.
Segmental Duplication Long stretches of DNA (usually over 1 kilobase) that are nearly identical and appear in multiple locations within a genome. They are major drivers of genome evolution and can predispose regions to further rearrangement via erroneous recombination.
Selective Pressure Any external factor (environmental, social, or biological) that affects the survival or reproductive success in a population. Individuals better suited to the pressure are more likely to survive and reproduce, driving natural selection and adaptation.
Splicing A crucial step in the process of mRNA maturation whereby non-coding regions (introns) are excised to produce a mature mRNA molecule containing the coding regions (exons).
Transcript An RNA molecule (such as mRNA, tRNA, or rRNA) produced through the transcription of a DNA template. It represents the first step in the conversion of genetic information into a functional product.
Transcription The process by which the information in a strand of DNA is copied into a molecule of RNA. This process is catalyzed by enzymes called RNA polymerases.
References
Aguileta G, Bielawski JP, Yang Z. Gene conversion and functional divergence in the beta-globin gene family. J Mol Evol. 2004 Aug.59(2):177-89. doi: 10.1007/s00239-004-2612-0.
Antonacci F, Dennis MY, Huddleston J, et al. Palindromic GOLGA8 core duplicons promote chromosome 15q13.3 microdeletion and evolutionary instability. Nat Genet. 2014 Dec.46(12):1293-302. doi: 10.1038/ng.3120.
Bailey JA, Eichler EE. Primate segmental duplications: crucibles of evolution, diversity and disease. Nat Rev Genet. 2006 Jul.7(7):552-64. doi: 10.1038/nrg1895
Betrán E, Long M. Expansion of genome coding regions by acquisition of new genes. Genetica. 2002. 115(1):65-80. doi: 10.1023/a:1016024131097
Betrán E, Wang W, Jin L, et al. Evolution of the phosphoglycerate mutase processed gene in human and chimpanzee revealing the origin of a new primate gene. Mol Biol Evol. 2002 May.19(5):654-63. doi: 10.1093/oxfordjournals.molbev.a004124.
Bizzotto S, Walsh CA. Making a Notch in the Evolution of the Human Cortex. Dev Cell. 2018 Jun 4.45(5):548-550. doi: 10.1016/j.devcel.2018.05.015.
Cao L, Peng B, Yao L, et al. The ancient function of RB-E2F pathway: insights from its evolutionary history. Biol Direct. 2010 Sep 20:5:55. doi: 10.1186/1745-6150-5-55.
Casola C, Betrán E. The Genomic Impact of Gene Retrocopies: What Have We Learned from Comparative Genomics, Population Genomics, and Transcriptomic Analyses? Genome Biol. Evol. 2017. 9(6):1351–1373. doi:10.1093/gbe/evx081
Charrier C, Joshi K, Coutinho-Budd J, et al. Inhibition of SRGAP2 function by its human-specific paralogs induces neoteny during spine maturation. Cell. 2012 May 11.149(4):923-35. doi: 10.1016/j.cell.2012.03.034.
Conrad B, Antonarakis SE. Gene duplication: a drive for phenotypic diversity and cause of human disease. Annu Rev Genomics Hum Genet. 2007:8:17-35. doi: 10.1146/annurev.genom.8.021307.110233.
Demuth JP, Hahn MW. The life and death of gene families. Bioessays 2009. 31: 29–39
Dennis MY, Nuttle X, Sudmant PH, et al. Evolution of human-specific neural SRGAP2 genes by incomplete segmental duplication. Cell. 2012 May 11.149(4):912-22. doi: 10.1016/j.cell.2012.03.033.
Dennis MY, Eichler EE. Human adaptation and evolution by segmental duplication. Curr Opin Genet Dev. 2016 Dec.41:44-52. doi: 10.1016/j.gde.2016.08.001.
Dornburg A, Mallik R, Wang Z. et al. Placing human gene families into their evolutionary context. Hum Genomics 16, 56 (2022). https://doi.org/10.1186/s40246-022-00429-5
Emanuel BS, Shaikh TH. Segmental duplications: an 'expanding' role in genomic instability and disease. Nat Rev Genet. 2001 Oct.2(10):791-800. doi: 10.1038/35093500.
Emes RD, Goodstadt L, Winter EE, et al. Comparison of the genomes of human and mouse lays the foundation of genome zoology. Hum Mol Genet 2003. 12: 701–709.
Esnault C, Maestre J, Heidmann T. Human LINE retrotransposons generate processed pseudogenes. Nat Genet. 2000. 24(4):363-7. doi: 10.1038/74184.
Fair T, Pollen AA. Genetic architecture of human brain evolution. Curr Opin Neurobiol. 2023 Jun.80:102710. doi: 10.1016/j.conb.2023.102710.
Feng Q, Moran JV, Kazazian Jr HH, et al. Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell. 1996. 29.87(5):905-16. doi: 10.1016/s0092-8674(00)81997-2.
Fiddes IT, Lodewijk GA, Mooring M, et al. Human-Specific NOTCH2NL Genes Affect Notch Signaling and Cortical Neurogenesis. Cell. 2018 May 31.173(6):1356-1369.e22. doi: 10.1016/j.cell.2018.03.051.
Florio M, Albert M, Taverna E, et al. Human-specific gene ARHGAP11B promotes basal progenitor amplification and neocortex expansion. Science. 2015 Mar 27.347(6229):1465-70. doi: 10.1126/science.aaa1975
Force A, Lynch M, Pickett FB, et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics. 1999 Apr.151(4):1531-45. doi: 10.1093/genetics/151.4.1531.
Fraser CM, Gocayne JD, White O, et al. The minimal gene complement of Mycoplasma genitalium. Science. 1995 Oct 20.270(5235):397-403. doi: 10.1126/science.270.5235.397.
Hahn MW, Demuth JP, Han SG. Accelerated rate of gene gain and loss in primates. Genetics. 2007a Nov.177(3):1941-9. doi: 10.1534/genetics.107.080077. Epub 2007 Oct 18.
Hahn MW, Han MV, Han SG. Gene family evolution across 12 Drosophila genomes. PLoS Genet. 2007b Nov.3(11):e197. doi: 10.1371/journal.pgen.0030197.
Hillier LW, International Chicken Genome Sequencing Consortium. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432, 695–716 (2004). https://doi.org/10.1038/nature03154
Hoffmann FG, Storz JF, Gorr TA, et al. Lineage-specific patterns of functional diversification in the alpha- and beta-globin gene families of tetrapod vertebrates. Mol Biol Evol. 2010 May.27(5):1126-38. doi: 10.1093/molbev/msp325. Epub 2010 Jan 4.
Iglesias-Ara A, Zubiaga AM. The stress of coping with E2F loss. Mol Cell Oncol. 2015 May 26.3(1):e1038423. doi: 10.1080/23723556.2015.1038423.
Iyer LM, Anantharaman V, Wolf MY, et al. Comparative genomics of transcription factors and chromatin proteins in parasitic protists and other eukaryotes. Int J Parasitol. 2008 Jan.38(1):1-31. doi: 10.1016/j.ijpara.2007.07.018. Epub 2007 Sep 15.
Juan D, Santpere G, Kelley JL, et al. Current advances in primate genomics: novel approaches for understanding evolution and disease. Nat Rev Genet. 2023 May.24(5):314-331. doi: 10.1038/s41576-022-00554-w.
Kaessmann H, Vinckenbosch N, Long M. RNA-based gene duplication: mechanistic and evolutionary insights. Nat Rev Genet. 2009 Jan.10(1):19-31. doi: 10.1038/nrg2487.
Kaessmann H. Origins, evolution, and phenotypic impact of new genes. Genome Res. 2010 20(10):1313-26. doi: 10.1101/gr.101386.109.
Kent LN, Leone G. The broken cycle: E2F dysfunction in cancer. Nat Rev Cancer. 2019 Jun.19(6):326-338. doi: 10.1038/s41568-019-0143-7.
Kimura, M. The Neutral Theory of Molecular Evolution. (Cambridge Univ. Press, Cambridge, (1983).
Lander ES, International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001). https://doi.org/10.1038/35057062
Liao X, Zhu W, Zhou J. et al. Repetitive DNA sequence detection and its role in the human genome. Commun Biol 6, 954 (2023). https://doi.org/10.1038/s42003-023-05322-y
Long M, Betrán E, Thornton K, et al. The origin of new genes: glimpses from the young and old. Nat Rev Genet. 2003. 4(11):865-75. doi: 10.1038/nrg1204.
Long M, VanKuren NW, Chen S et al.. New gene evolution: little did we know. Review Annu Rev Genet. 2013:47:307-33. doi: 10.1146/annurev-genet-111212-133301. Epub 2013 Sep 13.
Lupski JR, Wise CA, Kuwano A, et al. Gene dosage is a mechanism for Charcot-Marie-Tooth disease type 1A. Nat Genet. 1992 Apr.1(1):29-33. doi: 10.1038/ng0492-29.
Lynch M, Conery JS. The Evolutionary Fate and Consequences of Duplicate Genes. Science. 2000, 290(5494): 1151-1155. DOI: 10.1126/science.290.5494.1151
Marques-Bonet T, Kidd JM, Ventura M, et al. A burst of segmental duplications in the genome of the African great ape ancestor. Nature. 2009 Feb 12.457(7231):877-81. doi: 10.1038/nature07744.
McCarrey JR. Molecular evolution of the human Pgk-2 retroposon. Nucleic Acids Res. 18, 949–955 (1990).
McDonald-McGinn DM, Sullivan KE, Marino B, et al. 22q11.2 deletion syndrome. Nat Rev Dis Primers. 2015 Nov 19.1:15071. doi: 10.1038/nrdp.2015.71
Moody ERR, Álvarez-Carretero S, Mahendrarajah TA, et al. The nature of the last universal common ancestor and its impact on the early Earth system. Nat Ecol Evol 2024 Sep.8(9):1654-1666. doi: 10.1038/s41559-024-02461-1
Nadeau JH, Sankoff D. Comparable rates of gene loss and functional divergence after genome duplications early in vertebrate evolution. Genetics. 1997 Nov.147(3):1259-66. doi: 10.1093/genetics/147.3.1259.
Ohno S. Evolution by Gene Duplication. Springer-Verlag, Berlin Heidelberg. (1973).
Olender T, Jones TEM, Bruford E, et al. A unified nomenclature for vertebrate olfactory receptors. BMC Evol Biol. 2020 Apr 15.20(1):42. doi: 10.1186/s12862-020-01607-6.
Parker HG, VonHoldt BM, Quignon P, et al. An expressed fgf4 retrogene is associated with breed-defining chondrodysplasia in domestic dogs. Science. 2009 Aug 21.325(5943):995-8. doi: 10.1126/science.1173275.
Patthy L. Exon Shuffling Played a Decisive Role in the Evolution of the Genetic Toolkit for the Multicellular Body Plan of Metazoa. Genes (Basel). 2021 8.12(3):382. doi: 10.3390/genes12030382.
Pei B, Sisu C, Frankish A, et al. The GENCODE pseudogene resource. Genome Biology. 2012. 13 (9) R51. doi:10.1186/gb-2012-13-9-r51.
Prince VE, Pickett FB. Splitting pairs: the diverging fates of duplicated genes. Nat Rev Genet. 2002 Nov.3(11):827-37. doi: 10.1038/nrg928.
Schienman JE, Holt RA, Auerbach MR, et al. Duplication and divergence of 2 distinct pancreatic ribonuclease genes in leaf-eating African and Asian colobine monkeys. Mol Biol Evol. 2006 Aug.23(8):1465-79. doi: 10.1093/molbev/msl025. Epub 2006 Jun 2.
Schmutz J, Cannon SB, Schlueter J, et al. Genome sequence of the palaeopolyploid soybean. Nature 463, 178–183 (2010). https://doi.org/10.1038/nature08670
Sebat J, Lakshmi B, Troge J, et al. Large-scale copy number polymorphism in the human genome. Science. 2004 Jul 23.305(5683):525-8. doi: 10.1126/science.1098918.
Soto DC, Uribe-Salazar JM, Shew CJ, et al. Genomic structural variation: A complex but important driver of human evolution. Am J Biol Anthropol. 2023 Aug.181 Suppl 76(Suppl 76):118-144. doi: 10.1002/ajpa.24713.
Suzuki IK, Gacquer D, Van Heurck R, et al. Human-Specific NOTCH2NL Genes Expand Cortical Neurogenesis through Delta/Notch Regulation. Cell. 2018 May 31.173(6):1370-1384.e16. doi: 10.1016/j.cell.2018.03.067.
Suzuki IK, Vanderhaeghen P. Evolving Brains with New Genes. Opera Med Physiol 2018 Vol. 4 (3-4): 78-85. doi:10.20388/omp2018.003.0061
Templeton TJ, Iyer LM, Anantharaman V, et al. Comparative analysis of apicomplexa and genomic diversity in eukaryotes. Genome Res. 2004 Sep.14(9):1686-95. doi: 10.1101/gr.2615304.
Venter JC et al. The sequence of the human genome. Science. 2001, 291(5507):1304-51. doi: 10.1126/science.1058040.
Vervoort L, Vermeesch JR. The 22q11.2 Low Copy Repeats. Genes (Basel). 2022 Nov 11.13(11):2101. doi: 10.3390/genes13112101.
Vinckenbosch N, Dupanloup I, Kaessmann H. Evolutionary fate of retroposed gene copies in the human genome. Proc Natl Acad Sci U S A. 2006 28.103(9):3220-5. doi: 10.1073/pnas.0511307103.
Vollger MR, Guitart X, Dishuck PC, et al. Segmental duplications and their variation in a complete human genome. Science. 2022 Apr.376(6588):eabj6965. doi: 10.1126/science.abj6965.
Wang X, Gao B, Zhu S. Exon Shuffling and Origin of Scorpion Venom Biodiversity. Toxins (Basel). 2016 Dec 26.9(1):10. doi: 10.3390/toxins9010010
Wright JC, Mudge J, Weisser H, et al. Improving GENCODE reference gene annotation using a high-stringency proteogenomics workflow. Nat Commun. 2016. 7 11778. doi:10.1038/ncomms11778.
Xia S, Chen J, Arsala D, et al. Functional innovation through new genes as a general evolutionary process. Nat Genet. 2025 Feb.57(2):295-309. doi: 10.1038/s41588-024-02059-0.
Yilmaz F, Karageorgiou C, Kim K, et al. Reconstruction of the human amylase locus reveals ancient duplications seeding modern-day variation. Science. 2024 Nov 22.386(6724):eadn0609. doi: 10.1126/science.adn0609.
Zhang J, Zhang YP, Rosenberg HF. Adaptive evolution of a duplicated pancreatic ribonuclease gene in a leaf-eating monkey. Nat Genet. 2002 Apr.30(4):411-5. doi: 10.1038/ng852. Epub 2002 Mar 4.
Zhang J, Antony M. Dean AM, et al. Evolving protein functional diversity in new genes of Drosophila Proc Natl Acad Sci U S A. 2004. 16.101(46):16246-50. doi: 10.1073/pnas.0407066101.
Zhang J. Parallel adaptive origins of digestive RNases in Asian and African leaf monkeys. Nat Genet. 2006 Jul.38(7):819-23. doi: 10.1038/ng1812. Epub 2006 Jun 11.
Zhang YW, Liu S, Zhang X, et al. A functional mouse retroposed gene Rps23r1 reduces Alzheimer's beta-amyloid levels and tau phosphorylation. Neuron. 2009.64:328–340. doi: 10.1016/j.neuron.2009.08.036.
Zhou Q, Zhang G, Zhang Y, et al. On the origin of new genes in Drosophila. Genome Research, (2008) 18, 1446-1455. doi:10.1101/gr.076588.108
Zhou Q, Wang W. On the origin and evolution of new genes--a genomic and experimental perspective. J Genet Genomics. 2008 Nov.35(11):639-48. doi: 10.1016/S1673-8527(08)60085-5.
COMPARTIR